Quick Start¶
Basic usage¶
from densitree import SPADE
# X is any (n_cells, n_features) array
spade = SPADE(n_clusters=30, downsample_target=0.1, random_state=42)
labels = spade.fit_predict(X)
With a pandas DataFrame¶
Column names are automatically preserved in the result:
import pandas as pd
from densitree import SPADE
df = pd.read_csv("cytometry_data.csv")
markers = ["CD3", "CD4", "CD8", "CD19", "CD56"]
spade = SPADE(n_clusters=30, downsample_target=0.1, random_state=42)
spade.fit(df[markers])
# Cluster stats include median_CD3, median_CD4, etc.
print(spade.result_.cluster_stats_)
Visualization¶
# Static matplotlib plot
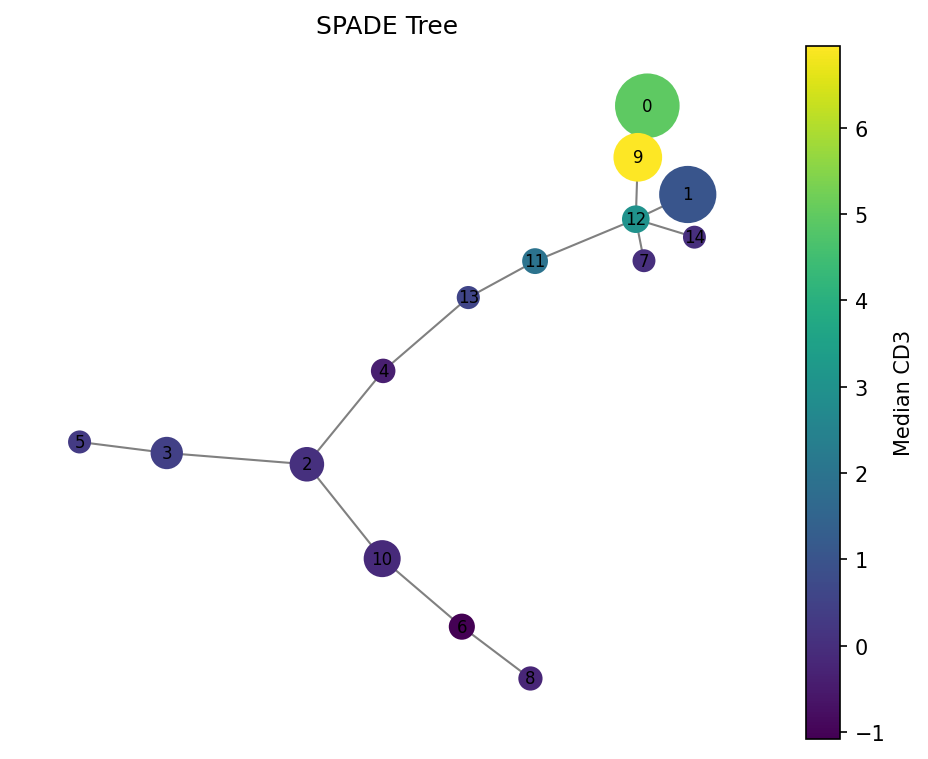
fig = spade.result_.plot_tree(color_by="CD3", backend="matplotlib")
fig.savefig("tree.png", dpi=150)
# Interactive plotly plot
fig = spade.result_.plot_tree(color_by="CD3", backend="plotly")
fig.show()
Choosing parameters¶
| Parameter | Default | Guidance |
|---|---|---|
n_clusters |
50 | 20--100 depending on expected complexity. More clusters = finer resolution but noisier tree. |
downsample_target |
0.05 | Fraction of cells retained. Lower = faster but may lose structure. 0.05--0.2 typical. |
knn |
5 | Neighborhood size for density estimation. 5--10 for most datasets. |
transform |
"arcsinh" |
Use "arcsinh" for cytometry (cofactor=5 for CyTOF, 150 for flow), "log" for counts, None if pre-transformed. |
cofactor |
150.0 | Arcsinh denominator. 5.0 for CyTOF, 150.0 for fluorescence. |